 
Mouse testes are homogenized and lysed in Sample preparation, and polysome RNAs are enriched by sucrose gradient centrifugation and used to calculate translation efficiency in Sample analysis.Ĭell populations and tissues exhibit unique gene expression profiles, which allow for characterizing and distinguishing cellular subtypes. Schematic illustrating the experimental design for polysome profiling in mouse testes. High efficiency and robustness compared to ribo-seq.Omit RNase digestion and RNA recovery from gel.Quickly obtain polysome RNAs from testes.This protocol allows to quickly isolate translating mRNAs from testes and analyze the discrepancy of translational efficiency in mouse testes from different mouse lines. Briefly, mouse testes are gently homogenized to release polysomes containing translating mRNAs, following polysome-bound mRNAs isolated by sucrose density gradient purification and characterized by RNA-seq. Here, we describe a protocol to identify translating mRNAs using polysome profiling. 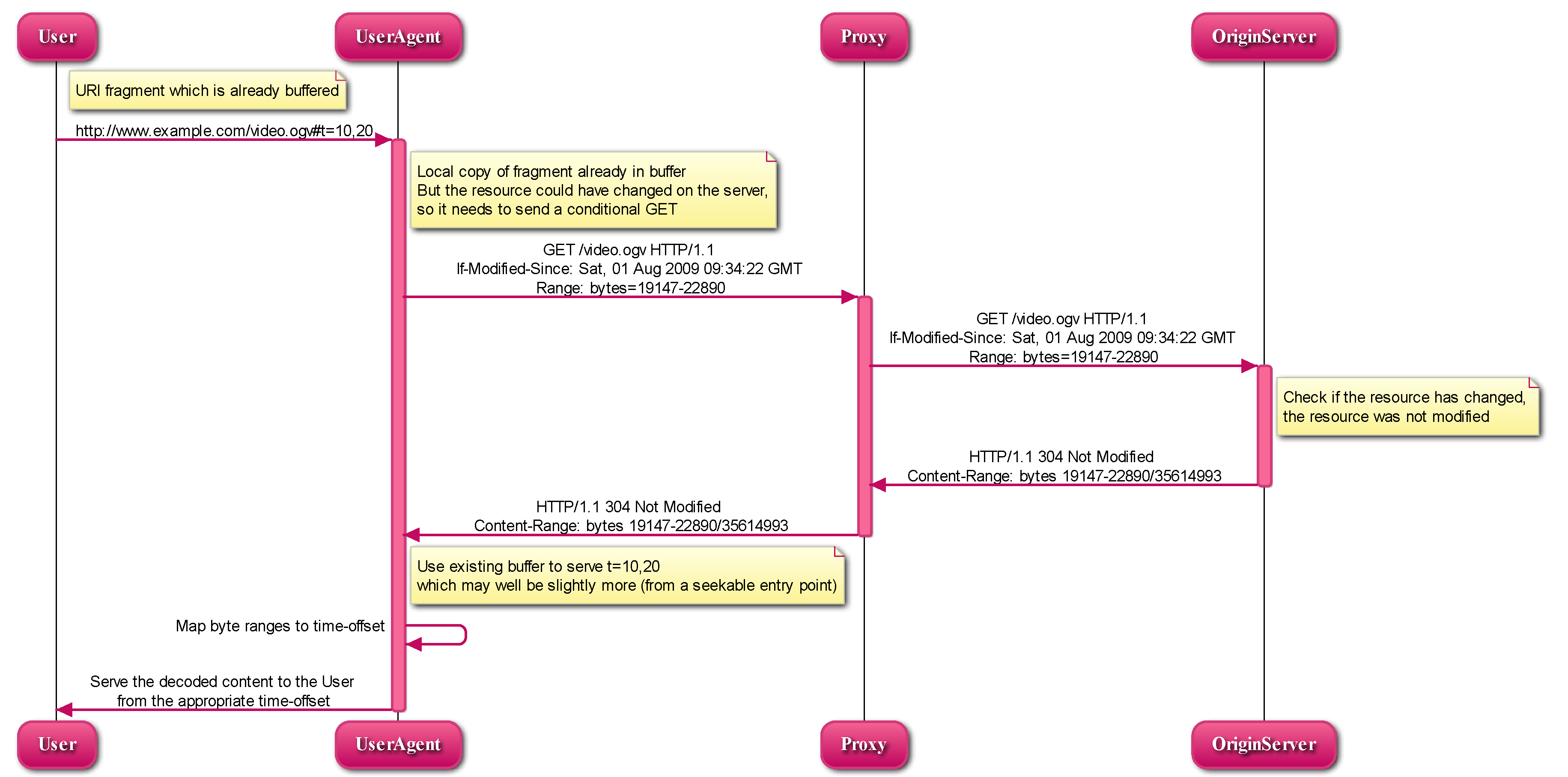
To understand the translation regulation during spermiogenesis, an overview of translational state of spermiogenic mRNAs is required. Spermiogenesis, i.e., the post-meiotic phase of male germ cell development, is a highly coordinated developmental process in which transcription and translation are decoupled because of nuclear condensation, resulting in translation regulation as the major mode for the regulation of gene expression in post-meiotic spermatids. 
Compared to ribosome profiling and translating ribosome affinity purification, polysome profiling is simpler and less time consuming in sample preparation and library constructions. Polysome profiling is widely used to isolate and analyze polysome fractions, which consist of actively translating mRNAs and ribosomes. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
